Within a month of the first novel coronavirus case reported in Wuhan, the 29,903 letters that make up the disease's genome were published online.
Now, researchers around the world are scrutinising the sequence, and the more than 50 sequences extracted from infected cells, to better understand and develop a vaccine for the disease.
The race is on. Last week, the World Health Organisation declared the coronavirus epidemic an international public health emergency.
More than 14,000 people have been infected in more than 20 countries, 304 are dead and there are five confirmed cases in the UAE.
The Chinese government is under scrutiny from the international community for its early communication on the virus, from the first known cases in early December to a lockdown of the city of 11 million about seven weeks later.
But the speed with which Chinese researchers mapped the novel coronavirus’ genome was practically unprecedented – and that is worth noting, experts said.
"Our ability to sequence the genome of viruses and share them globally in an open-source fashion can significantly improve our effectiveness and speed in responding to viral outbreaks, thus reducing casualties," Nadine Nehme, chief science officer at Medicus, told The National.
During the Sars outbreak in 2002, it took scientists about 20 months to go from genetic sequencing to the first phase of human trials of a vaccine.
By that time, the outbreak was under control, but nearly 800 people had died.
The timeline for a vaccine is expected to be reduced to three months with the Wuhan virus as time, talent and capital are dedicated to tackling the epidemic, the US National Institute of Health said.
Two weeks after the sequence was first published, the Coalition for Epidemic Preparedness Innovations, a public-private investment group based in Norway, opened its coffers to invest $9 million into three programmes to advance vaccine candidates into clinical testing as quickly as possible.
It is aiming to go from the gene sequence, first published on January 10, to clinical testing of a vaccine in 16 weeks.
There are three main uses for mapping the virus' genome, Anamika Mehta, an applied molecular biology and clinical genetics specialist in Dubai, told The National.
First, the sequence helps in diagnosing the cause of disease when a person has suspicious symptoms such as coughing, a fever and shortness of breath.
A doctor facing a patient like this would ask, “Is this just a harmless common cold, or could it progress to fatal pneumonia?”, Ms Mehta said.
“Different viruses have differing letter sequences and determining the specific viral sequence in the affected person helps pinpoint the cause,” she said.
“Diagnostic tests based on this viral sequence are now being used to confirm those affected specifically with the Wuhan virus.”
Second, comparing the sequence of this new virus with those from previous epidemics “can give us a hint of what to expect”, Ms Mehta said.
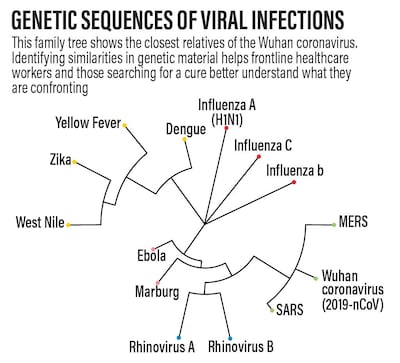
Mapping the Wuhan virus allowed researchers to place it in a "family tree".
Similarities in genetic material showed it was a “close cousin of the viruses responsible for two other well-known epidemic outbreaks: the Sars virus from the 2003 epidemic in China, and the Mers virus from the 2012 epidemic in Saudi Arabia”, she said.
The similarity between the three sequences is about 80 per cent so all three viruses have a fairly recent origin in humans and can be expected to have similar symptoms.
“Interestingly, all three are more distant cousins of the harmless common cold,” Ms Mehta said.
“The third use is to pinpoint the source of the virus: where was it hiding before it suddenly broke loose into humans in the seafood market in Wuhan more than a month ago?
"Which animal was the reservoir for this virus, and which one was intermediary in the transmission to humans?”
Passing the disease from one living thing to another causes viral infections to adapt and further confound immune systems.
Researchers will need to keep a close eye on new samples of the genome sequence to figure out how the virus adapts and changes as it infects more people.
As for where the technology is headed, Illumina, a biotechnology company in California, made the sequencer machine used to identify and publish the genomic profile of the coronavirus online.
Yagnessh Kesavan, a team leader at Genomics MEAA in Dubai, which distributes Illumina equipment, told The National that identifying an unknown organism by sequencing its genome can happen in less than 24 hours practically anywhere in the world.
“Along with a couple of suitcases of lab equipment” and a genome sequencer made by the company known as the iSeq, “we could have a travelling laboratory and set up a full-functioning sequencing facility hours from arrival in any part of the world by just plugging in the iSeq, running a systems check and beginning to process samples immediately", Mr Kesavan said.
More accurate diagnosis and treatment will lead to reduced hospital stays and the more considered use of antibiotics, resulting in an improved level of health care.
Ms Mehta said speed was important but the most important aspect of battling infectious diseases is a better understanding of the way a virus functions to overcome a body’s first line of defence: our immune system.
She believes the day will soon come when altering the body’s response to a virus, known as immunotherapy, will be precise and effective using nanoparticles, which are too tiny to be seen with an ordinary microscope.
“The Wuhan virus is killing people because their immune system cannot fight it," Ms Mehta said.
"But what if we could come up with immunotherapy strategies for such infection where a nanoparticle can teach our immune cells to kill these viruses with precision?”.